 
Retrotransposon probes Copia, Gypsy, Jockey, and RTE showed a more pulverized hybridization pattern, with the presence of small interstitial and/or terminal blocks. DNA transposon probes hAT, Helitron, and Tc1-mariner showed mostly scattered signals, with the presence of some blocks. The transposable elements that stood out in the transcriptome were selected for fluorescent in situ hybridization. Phylogenetic analyses were performed with the protein domains, evidencing differences between Tc1-mariner sequences, which may be related to possible horizontal transfer events. Among Class I elements or retrotransposons, most were characterized as LINE. Of the 367 transcripts identified as TEs, 202 belong to Class II, with emphasis on the TIR order. Through RNA-Seq and de novo assembly, 60,009 transcripts were obtained, which were annotated locally via Blastn against specific databases. heros transcriptome, evidencing their chromosomal distribution. Therefore, the objective of the current work was to identify the different groups of transposable elements present in the E. In the case of Euschistus heros, considered the main stink bug in the soybean crop in Brazil, little is known about the participation of these elements. 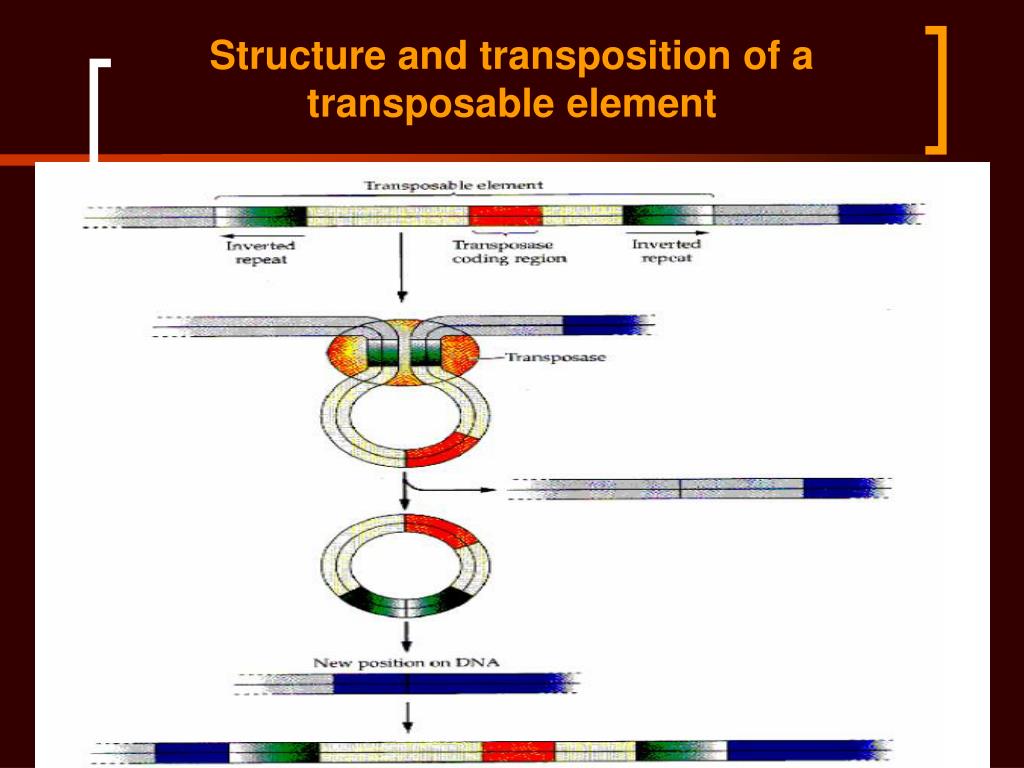
Their distribution is very dynamic among organisms, and despite advances, there are still gaps in the understanding of the diversity and evolution of TEs in many insect species. Kronmiller BA, Wise RP (2008) TEnest: automated chronological annotation and visualization of nested plant transposable elements.Transposable elements (TEs) are DNA sequences capable of moving within the genome. Springer, Cham, pp 113–130Īnderson SN, Stitzer MC, Brohammer AB, Zhou P, Noshay JM, O’Connor CH, Hirsch CD, Ross-Ibarra J, Hirsch CN, Springer NM (2019) Transposable elements contribute to dynamic genome content in maize. In: Origin and evolution of biodiversity. Su W, Sharma SP, Peterson T (2018) Evolutionary impacts of alternative transposition. Szuplewska M, Czarnecki J, Bartosik D (2014) Autonomous and non-autonomous Tn. 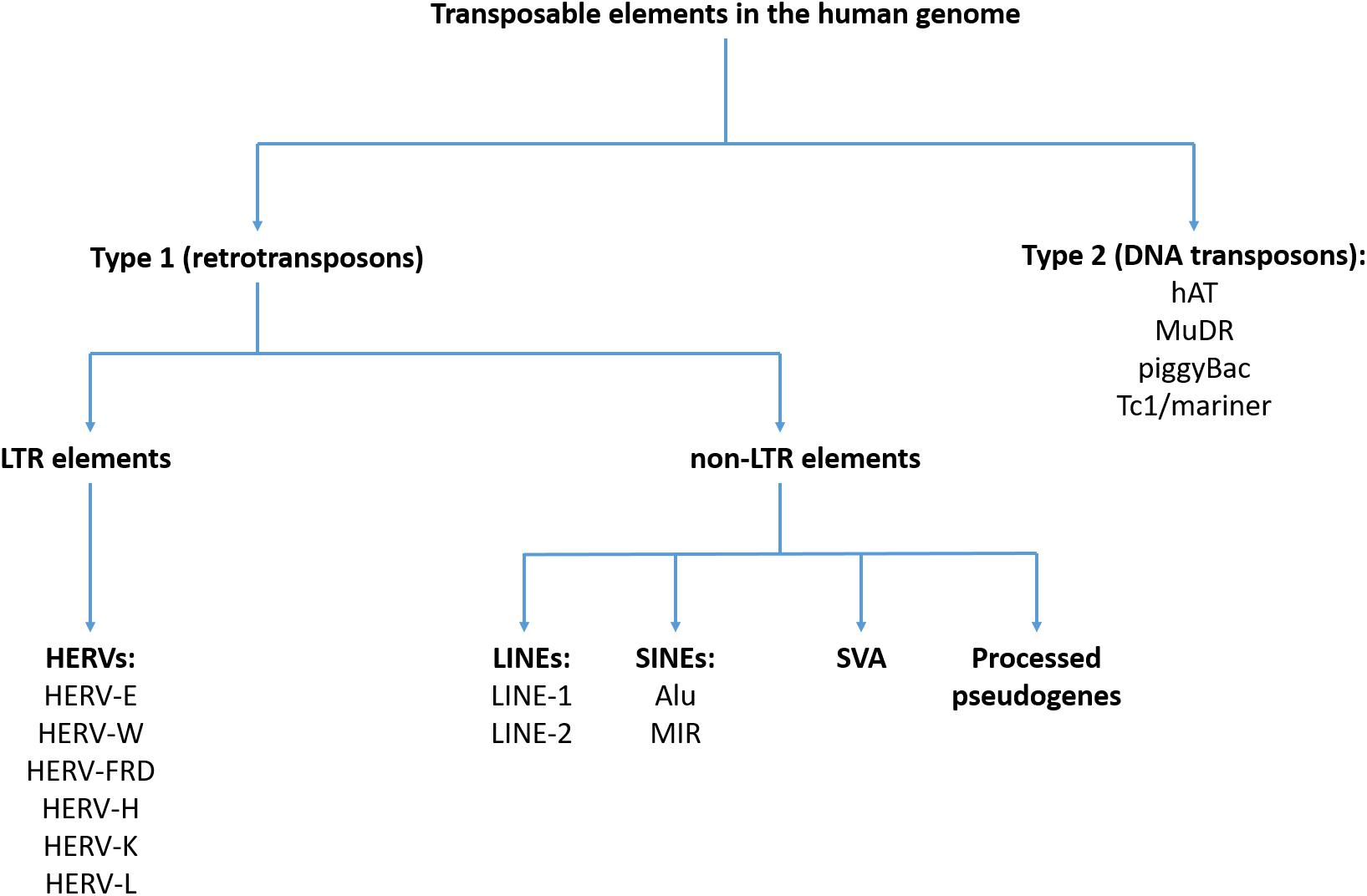
Pray LA (2008) Transposons: the jumping genes. Xiong W, He L, Lai J, Dooner HK, Du C (2014) HelitronScanner uncovers a large overlooked cache of Helitron transposons in many plant genomes. Su W, Gu X, Peterson T (2019) TIR-learner, a new ensemble method for TIR transposable element annotation, provides evidence for abundant new transposable elements in the maize genome. Shi J, Liang C (2019) Generic repeat finder: a high-sensitivity tool for genome-wide de novo repeat detection. Ou S, Jiang N (2018) LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Ou S, Jiang N (2019) LTR_FINDER_parallel: parallelization of LTR_FINDER enabling rapid identification of long terminal repeat retrotransposons. Įllinghaus D, Kurtz S, Willhoeft U (2008) LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. Ou S, Su W, Liao Y, Chougule K, Agda JRA, Hellinga AJ, Lugo CSB, Elliott TA, Ware D, Peterson T, Jiang N, Hirsch CN, Hufford MB (2019) Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Lerat E (2010) Identifying repeats and transposable elements in sequenced genomes: how to find your way through the dense forest of programs. Goerner-Potvin P, Bourque G (2018) Computational tools to unmask transposable elements. Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O et al (2007) A unified classification system for eukaryotic transposable elements. Science 326:1112–1115įeschotte C, Jiang N, Wessler SR (2002) Plant transposable elements: where genetics meets genomics. Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA (2009) The B73 maize genome: complexity, diversity, and dynamics. McClintock B (1950) The origin and behavior of mutable loci in maize. McClintock B (1948) Mutable loci in maize. Here, we present an overview of the EDTA pipeline and a detailed manual for its use. The selected programs are evaluated by their ability to identify true TEs as well as to exclude false candidates. The development of EDTA is based on the benchmarking results of a collection of TE annotation methods. In this chapter, we introduce EDTA (Extensive de novo TE Annotator), a new comprehensive pipeline composed of high-quality tools to identify and annotate all types of TEs. Each of these tools has advantages and disadvantages. These computational tools can be categorized into three types based on their underlying approach: homology-based, structural-based, and de novo methods. With the growth of sequencing technologies, various computational pipelines and software programs have been developed to facilitate TE identification and annotation. 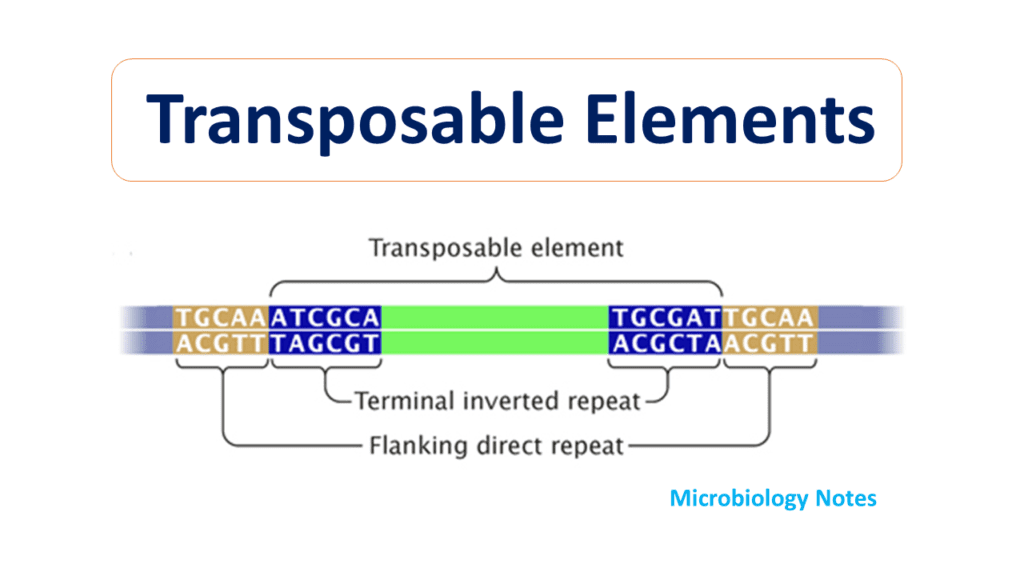
Transposable elements (TEs) are important contributors to genome structure and evolution. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
